
A web server for scaffolding contigs based on exemplar breakpoint distance

A web server for scaffolding contigs based on exemplar breakpoint distance
EBD-scaffolder is a web server that allows users to conveniently and accurately scaffold (i.e., order and orient) the contigs of a target draft genome using a single reference genome from a related organism. The scaffolding function of EBD-scaffolder was implemented using an integer linear programming algorithm that we specifically designed to solve the exemplar breakpoint distance (EBD) based scaffolding problem. The web server of EBD-scaffolder is free and open to all users with no login required.
EBD-scaffolder provides an easy-to-use web interface (see Figure 1 for an example). For convenience, users can either select one of the examples (1) we have already prepared for testing or running EBD-scaffolder, or submit a job by following the procedures described below.
Users can click the "Help" tab page (7) to view instructions on how to run EBD-scaffolder.
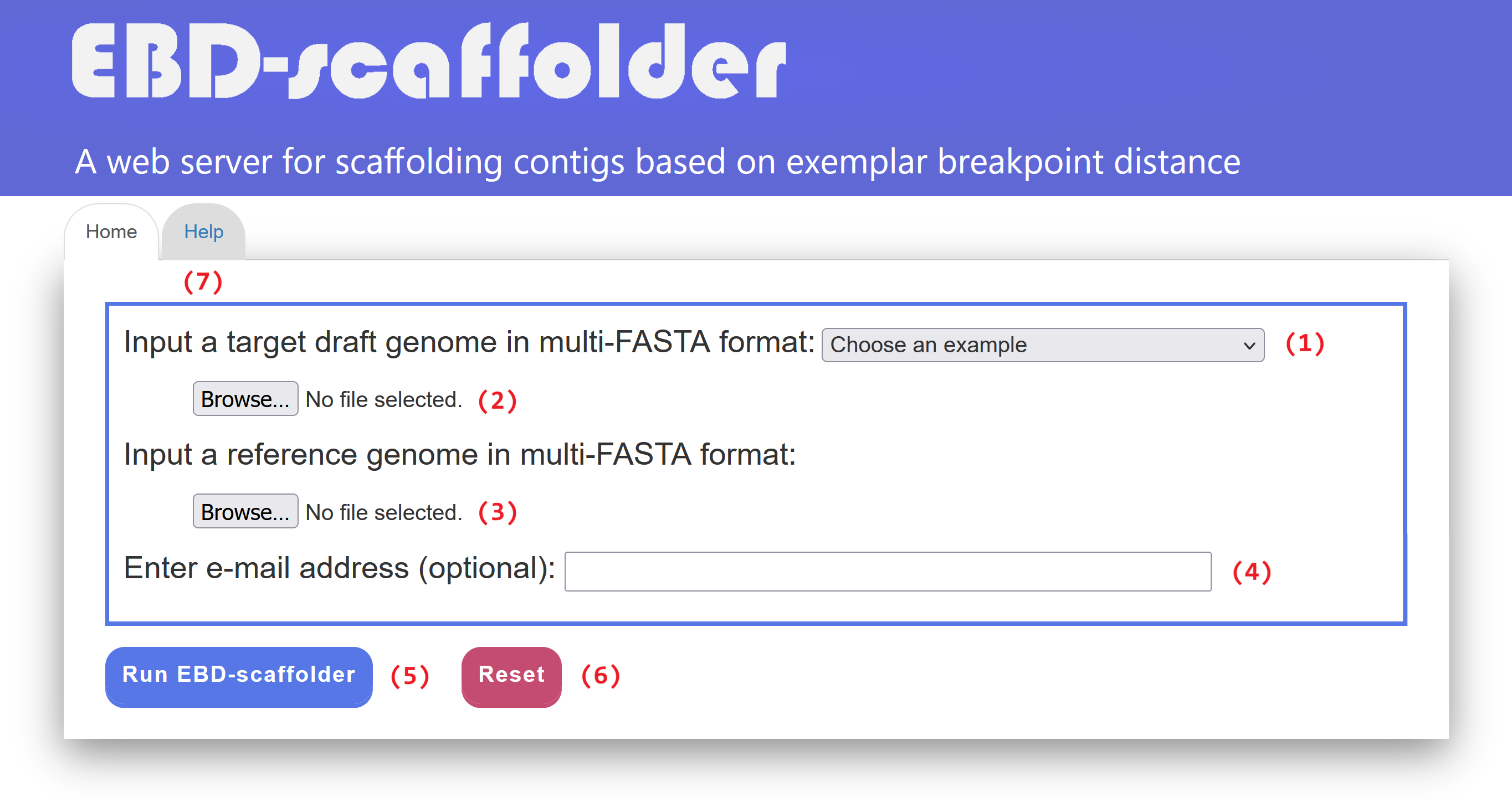
Figure 1: Web interface of EBD-scaffolder.
EBD-scaffolder outputs its scaffolding results across four tab pages: (1) Input data & parameters, (2) Circos plot validation, (3) Dotplot validation, and (4) Scaffolds of target.
In the "Input data & parameters" tab page (see Figure 2 for an example), EBD-scaffolder provides links to display the sequence data of the user-inputted target and reference genomes, as well as links to show the IDs of their contigs and scaffolds. The format of IDs begins with a three-letter prefix (e.g., CTG for contigs or SCF for scaffolds), followed by an underscore (_) and at least one digit (e.g., CTG_1 for a contig and SCF_1 for a scaffold). In addition, the EBD-scaffolder provides a dotplot, enabling users to visually inspect the markers identified by Sibelia between the input target and reference genomes prior to scaffolding. In the dotplot display, the input target genome is plotted along the y-axis, while the reference genome is plotted along the x-axis. Horizontal dashed lines separate the contigs within the target genome, and vertical dashed lines separate the scaffolds within the reference genome. In addition, forward and reverse markers are displayed as blue and red lines, respectively, with the beginning and end of each line represented by two unfilled points. It is important to note that the contigs in the target genome have not yet been scaffolded, resulting in the markers in the dotplot being scattered and disordered, as shown in Figure 2. Users have the option to sort the contigs of the input target genome by size using the toggle switch "Sort by contig size". They can also show or hide the IDs of contigs and scaffolds used in the EBD-scaffolder by clicking the toggle switch "Show contig/scaffold IDs". If needed, users can download the dotplot of the input target and reference genomes in scalar vector graphics (SVG) format by clicking the "Save as SVG file" button.
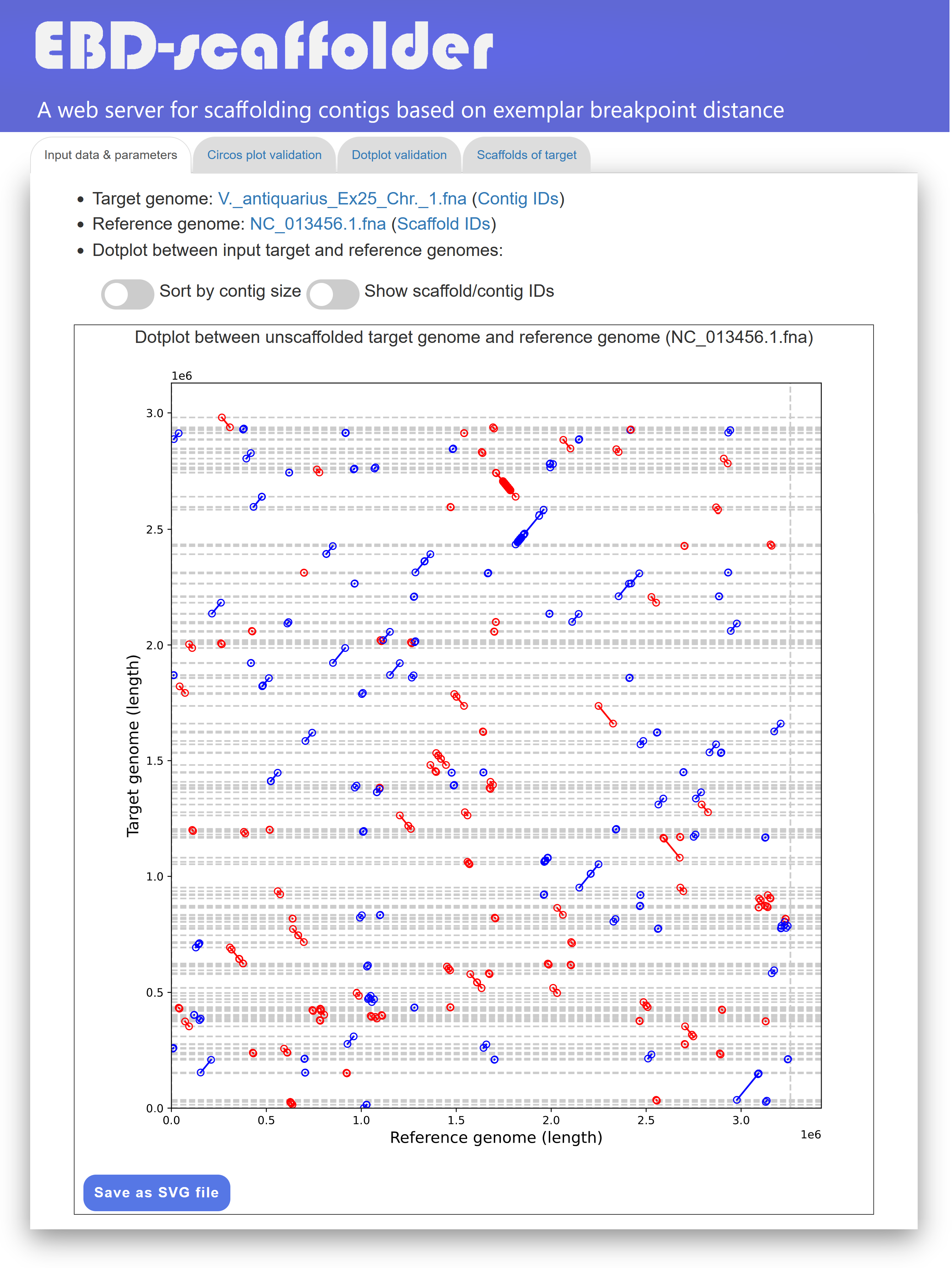
Figure 2: A display of the "Input data & parameters" tab page.
In the "Circos plot validation" tab page (see Figure 3 for an example), EBD-scaffolder displays its total running time, along with a Circos plot depicting the scaffolding result between the target and reference genomes. In this Circos plot, scaffolds from the target genome (blue) and the reference genome (green) are arranged along the outer circle. The markers between them are displayed in color and alternately placed on two layers of the inner circle based on their direction: forward markers on the outer layer and reverse markers on the inner layer. Additionally, identical markers shared by the target and reference genomes are connected via inner links within the inner circle. The number of crossing inner links in the Circos plot serves as an accuracy measure for the scaffolding result. Specifically, if the contigs of the target genome are well scaffolded in accordance with the reference genome, the number of crossing inner links between them should be low. As shown in Figure 3, the Circos plot provided by EBD-scaffolder offers a convenient and effective way for users to visually validate whether the contigs of the target genome are properly scaffolded with respect to the reference genome. In the Circos plot display, users can show the contig IDs of the target genome by toggling the switch labeled "Show all contig IDs". They can also download the Circos plot of the scaffolded target and reference genomes in SVG format by clicking the "Save as SVG file" button.
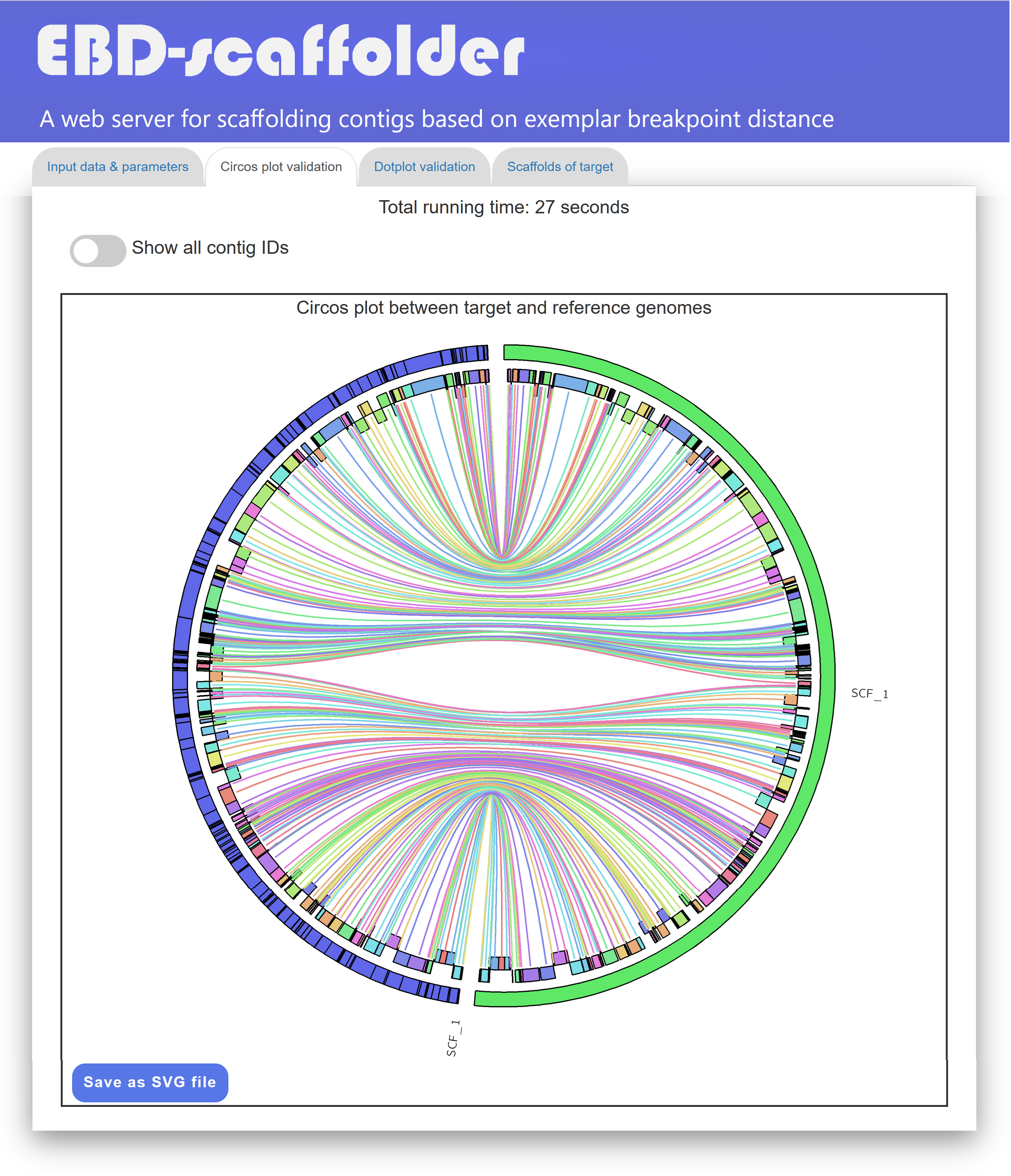
Figure 4: A display of a Circos plot between scaffolded target genome and reference genome.
In the "Dotplot validation" tab page (see Figure 4 for an example), EBD-scaffolder presents its scaffolding results using a dotplot to display the arragnement of identical markers between the scaffolded target genome and the reference genome. If the contigs of the target genome are perfectly scaffolded with respect to the reference genome, the markers displayed in the dotplot will extend from the lower-left corner to the upper-right corner (as shown in Figure 4) or from the upper-left corner to the lower-right corner. Such a dotplot display of the scaffolding result provides users with another convenient way to visually confirm whether the contigs of the target genome are properly scaffolded according to the reference genome.
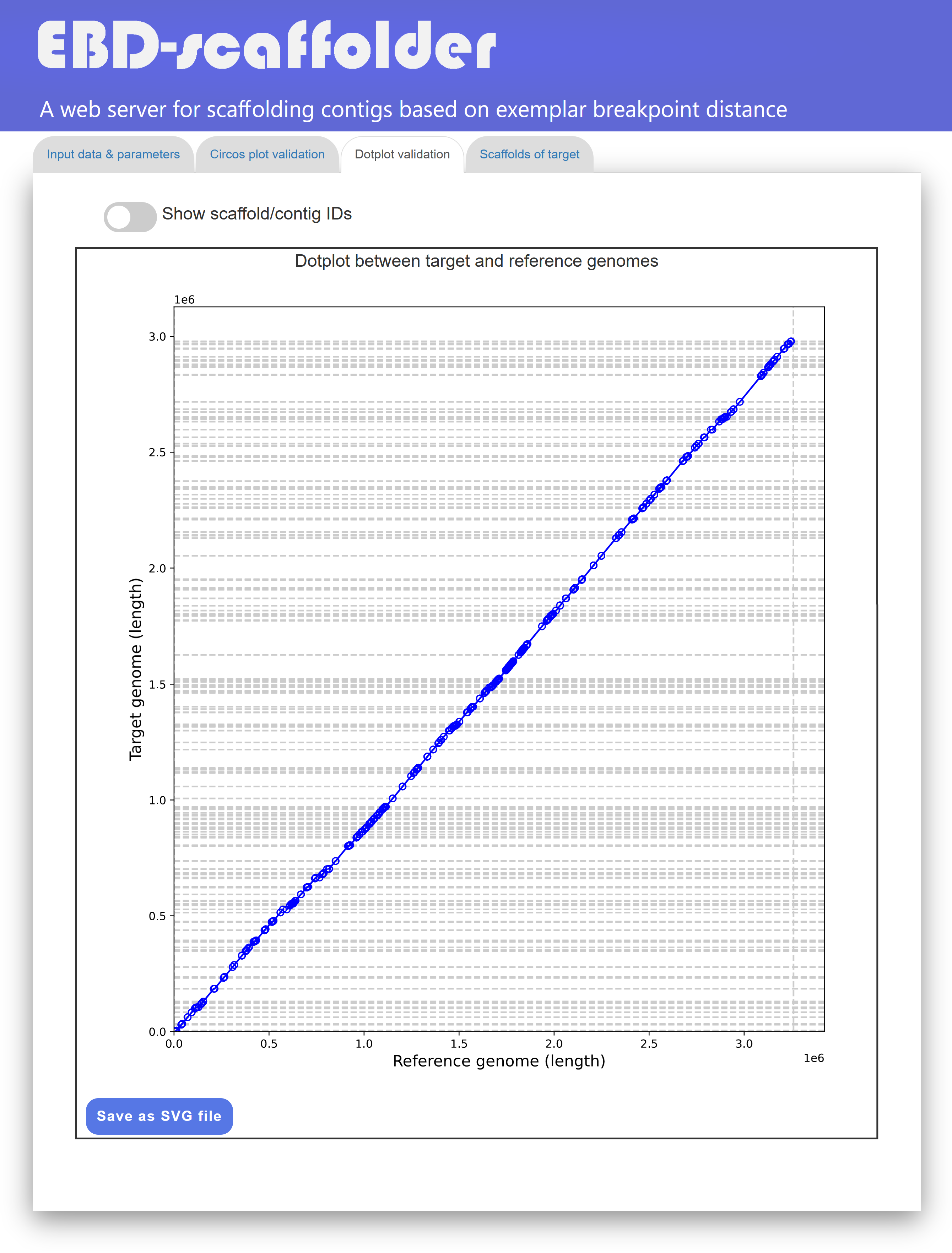
Figure 4: A display of the "Dotplot validation" tab page.
In the "Scaffolds of Target" tab page (see Figure 5 for an example), EBD-scaffolder presents its scaffolding result in a tabular format, allowing users to view the scaffolds of the target genome in detail. The scaffolds listed in the table are sorted by their sizes, which are calculated as the sum of the contig sizes. The contigs of each scaffold, along with their orientation (0 for forward and 1 for reverse), sequence and length, are organized in a table according to their order within the scaffold. For downstream analyses, users can either download the scaffolds of the target genome in a tab-delimited text format or in a comma-delimited CSV format by clicking the "Download scaffolds (.txt)" or "Download scaffolds (.csv)" button, respectively. In addition, users can download scaffold sequences in text format by clicking the `Download sequences' button, in which contig sequences within the same scaffold are separated by 100 Ns.
A standalone version of EBD-scaffolder can be downloaded as a single zipped file by clicking this download link.
You need to install the following prerequisite tools to properly run EBD-scaffolder and ensure that their paths are added to your PATH environment variable.
Run the following command to compile EBD-scaffolder. Note that if you download and use the new version of Gurobi, then you need to re-compile EBD-scaffolder first.
makeUse the following command to run EBD-scaffolder:
./EBD-scaffolder -t <TARGET GENOME PATH> -r <REFERENCE GENOME PATH> -o <OUTPUT DIRECTORY> -s <SIBELIA CONFIGURATION> -m [PARAMETER M FOR SIBELIA] -T [THREAD NUMBER] -i <TIME LIMIT>The scaffolding result of EBD-scaffolder in tabular format is stored in a file named "ScaffoldResult" at the "OUTPUT DIRECTORY" directory.
./EBD-scaffolder -t target.fna -r reference.fna -o Results -s parameter/paraset_bacterial -m 50 -T 16 -i 14400Corresponding author: Prof. Chin Lung Lu (Email: cllu@cs.nthu.edu.tw)